INTRODUCTION
As Westernized dietary habits and urban lifestyles become more commonly embraced, the incidence of colorectal carcinoma (CRC) is expected to spike in the near future. At present, though the age adjusted incidence rate of CRC is still low in developing countries, the lack of awareness and affordable healthcare facilities result in detection of the disease at an already advanced stage.
1,2 Overall, CRC is one of the most common causes of cancer-related mortality worldwide.
3 Thus, it is necessary to introduce effective population-based screening programs, especially in developing nations where such programs do not exist. However, one of the drawbacks of conventional colonoscopic screening is that it can only detect macroscopic mucosal lesions, such as adenomas, when the lesion has already undergone complex genetic changes. Hence, future screening techniques should aim to detect early microscopic preneoplastic lesions before the development of mass lesions has occurred. Currently, narrow band imaging, confocal endoscopes, and chromoendoscopy are increasingly available in major cities, which may enable the
in vivo detection of microscopic preneoplastic lesions, such as aberrant crypt foci (ACF).
4
ACF-like lesions were first identified during the microscopic examination of mutagen treated colonic mucosa in rodents and were subsequently identified in human colonic mucosae.
5,6,7,8 In a recently published study, we demonstrated the up-regulation of cancer stem cell (CSC) marker CD24 in ACF, similar to that in CRC, in comparison to that in the normal colon.
9 Kudo et al.
10 also showed the up-regulation of CSC marker LGR-5 in the ACF of mutagen-treated mice. Despite these interesting findings, the lack of a comprehensive understanding of the ACF-like lesions in human colon has not led to the clinical use of these lesions as markers of early disease in the human colonic mucosa. It remains unclear whether all ACF-like lesions in the human colon are potentially malignant, or if a specific morphological, topographic, or genetic subgroup is more prone to malignant change. In our previous study, we observed that ACF-like lesions could be identified in a wide variety of colectomy specimens from patients undergoing surgery because of various indications.
9 In this study, we sought to examine whether topographic, histological, and genetic changes differ in ACF identified in different clinical groups and also to identify the characteristics of high-risk ACF in the human colon.
METHODS
1. Patients Characteristics and Sample Preparation
This was a cross-sectional study conducted on a total of 270 fresh colorectal resection specimens, irrespective of the indication for surgery. Among these 270 specimens, ACF were identified in 107 specimens. Cases with a history of neoadjuvant therapy were excluded. The colectomy specimens with ACF were divided into following 3 clinical groups. Group A included colectomies performed for CRC (n=67). Group B (n=40) contained the disease controls and were further subdivided into group Bc (n=23; colectomies performed for conditions with an established link of colonic carcinoma development; e.g., 19 surgeries performed for chronic IBD of more than 10 years' duration, 4 adenomatous polyposis, etc.) and group Bn (n=17; colectomies performed for diseases having no known link with any neoplastic lesion development, e.g., 7 ischemic bowel disease, 3 tuberculosis, 2 intestinal perforation, 4 Hirschsprung disease, and 1 gastric gastro-intestinal stromal tumor).
After receiving the fresh specimens, macroscopically normal-looking colonic mucosa situated at least 10 cm away from the tumor was identified and dissected out from the underlying submucosa. For all specimens, the shaved mucosal flap size as 2.5 cm×2.5 cm with an area of approximately 6.25 cm2. These mucosal flaps were then stretched out on a paraffinized plate and fixed overnight in 10% neutral buffered formalin. In the non-tumor group, mucosal flaps were randomly sampled.
2. Identification and Immunohistochemistry of ACF
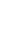
Following fixation, these flaps were stained with 0.5% methylene blue and observed for ACF at low-power magnification (×40 magnification) under an Olympus BX50 microscope (Tokyo, Japan). Single crypts or groups of distorted and darkly stained crypts were marked with water-resistant tissue paint (Davidson Marking System; Bradley Product, Inc., Bloomington, MN, USA). During the gross identification of ACF, various parameters such as elevation of the ACF, Kudo's pit pattern,
10 crypt pattern according to our previous descriptions, size, frequency, and multiplicity of the ACF were assessed and recorded.
9 Each case was photographed at ×40 magnification for computer-assisted image analysis. The marked areas of the flaps were then cut into thin strips and processed for paraffin embedding to confirm the ACF by H&E staining. Histological confirmation of ACF, along with mucosal histology, the presence or absence of dysplasia, and hypercrinia were noted by 2 pathologists independently. Cases with ACF confirmed on histology were then selected for the next part of the analysis. The subsequent part of analysis involved immunohistochemistry (IHC) and PCR for
KRAS and
BRAFV600E mutation detection. For molecular analysis, the blocks were stored separately in waterproof containers at 4℃.
IHC markers studied on the ACF-positive tissue sections included: (1) mismatch repair (
MMR) protein markers, MLH1, MSH2, PMS2, MSH6; and (2) marker of cell cycle regulation, p53. Tissue sections (5 µm) were cut from the paraffin blocks and deparaffinized. Endogenous peroxidase was blocked using 4% hydrogen peroxide, followed by antigen retrieval through boiling in citrate buffer. Primary monoclonal antibodies for MLH1 (1:400; Bio SB, Santa Barbara, CA, USA), PMS2 (1:200; Spring Bioscience Corp., Pleasanton, CA, USA), MSH2 (1:200, Bio SB), MSH6 (1:100, Bio SB), and p53 (1:400, Spring Bioscience) were applied on the sections, and the slides were incubated overnight at 4℃. The reaction product was developed with 3,3′-diaminobenzidine and counterstained with hematoxylin. Appropriate positive and negative controls were used. The IHC slides were then evaluated by comparing the staining pattern in the ACF with that in the adjacent normal-appearing crypts. IHC for MMR proteins was considered a surrogate marker for assessing microsatellite instability (MSI). Positive staining denoted functional expression of the gene product, and implied microsatellite stability. Any staining >10% of the epithelial cell nuclei was considered positive, i.e., microsatellite stable (MSS); staining in <10% of cells or negative staining was considered to represent MSI. p53 positivity in >10% of tumor cell nuclei was considered positive and to represent the surrogate marker for
TP53 mutation.
11 Expression of all the markers was noted separately in the ACF, tumors, and normal crypts.
3. Identification of KRAS and BRAF Mutations
KRAS mutation was tested in a total of 36 cases of ACF: CRC-associated ACF (n=26), ACF associated with control group Bc (n=4), and ACF associated with control group Bn (n=6). The cases were chosen based on presence of numerous ACF (>10 ACF/mm2) in the colonic mucosa. The representative areas were then microdissected out from unstained 10-µm sections (for a total of approximately 50 µg) and placed in lysing buffer until proper tissue homogenization occurred. Subsequent steps involved precipitating DNA with the help of a binding buffer (Purelink Genomic DNA Mini Kit, Life Technologies, Carlsbad, CA, USA), repeated washing in the buffers provided, and then elution using micro-column-based technique. The DNA content and quality were measured using a NanoDrop machine. Mutation-specific forward and reverse primers (5′-GAATATAAACTTGTGGTAGTTGGAGCT-3′ [F] and 5′-ATCGTCAAGGCACTCTTGCCTAC-3′ [R]) were used to target codons 12 and 13 of the KRAS gene and their flanking regions, with a calculated amplicon size of 56 bp. DNA extracted from the blood of normal subjects was used as a negative control (wild-type KRAS ). We performed high-resolution melting curve analysis using an AriaMX real-time PCR (qPCR) machine from Agilent Technologies (Santa Clara, CA, USA). The high-resolution melting curve analysis results were then validated by forward-strand sequencing in 10 representative samples (including the blood sample). For this part of the analysis, a separate primer was used to yield an amplicon of 254 base pairs, covering exon 2 of the KRAS gene. This amplicon was run by gel electrophoresis, eluted, and sequenced by Sanger sequencing. The results were analyzed using sequencing software by Applied Biosystems (Foster City, CA, USA).
BRAF mutation analysis was performed on 25 randomly selected cases from all 3 study groups. Extracted DNA was subjected to qPCR using
BRAF V600E-specific probes, as described in an earlier study.
12 Briefly, a 25-µL reaction was prepared using a Taqman PCR core reagent kit (Applied Biosystems) and 900-nM forward and reverse primers, 250 nM of a wild-type probe, and a mutation-specific probe. The Taqman-MGB probes were specific for the
V600E mutation (c.1799T>A) in BRAF exon 15. qPCR was performed using a Qiagen Rotor Gene-Q machine with the following conditions: 96℃ for 10 minutes, 40 cycles of 96℃ for 15 seconds, and 60℃ for 30 seconds with acquisition in carboxyfluorescein (FAM) (for mutation) and VIC fluorescent dye (for wild-type) channels. Analysis and mutation calls were made using Rotor Gene Q software.
4. Statistical Analysis
Data analysis was performed using Stata 12.1 software (College Station, TX, USA). Mean ACF number and density were compared between the study groups using Kruskal-Wallis ANOVA. ACF density with laterality was analyzed using the Student
t-test. ACF histology was analyzed in relation to the groups using the Fisher exact test. Association of hypercrinia with the study groups was performed using chi-square test. To evaluate the association of hypercrinia with laterality and ACF number simultaneously, the linear trend chi-square test was used. Analysis of Kudo et al.'s results
10 and those of our group using various parameters like laterality, ACF histology, and presence of hypercrinia was performed using the chi-square test. Similarly, all the IHC markers and mutation results were analyzed in relation to groups and parameters using the chi-square and linear trend chi-square tests. The study protocol was approved by the Institutional Review Board at the All India Institute of Medical Sciences, New Delhi, India (IRB numbers: IESC/T-34/03.01.2014, OT-3/30.09.2015) and performed in accordance with the principles of the Declaration of Helsinki. Informed consent was waived.
DISCUSSION
In our study, ACF-like lesions were identified in the grossly normal-looking colonic mucosa of patients with a variety of diseases including polyposis, adenomas and CRC, as well as in association with IBD, intestinal tuberculosis, inflammatory perforation, ischemia, and so forth.
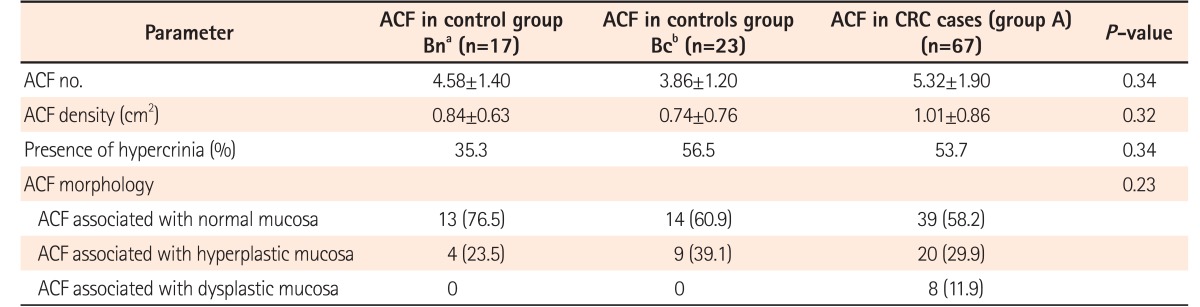
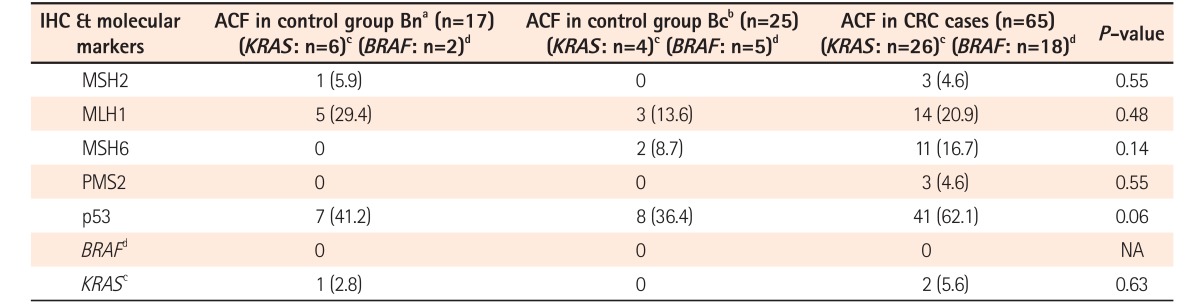
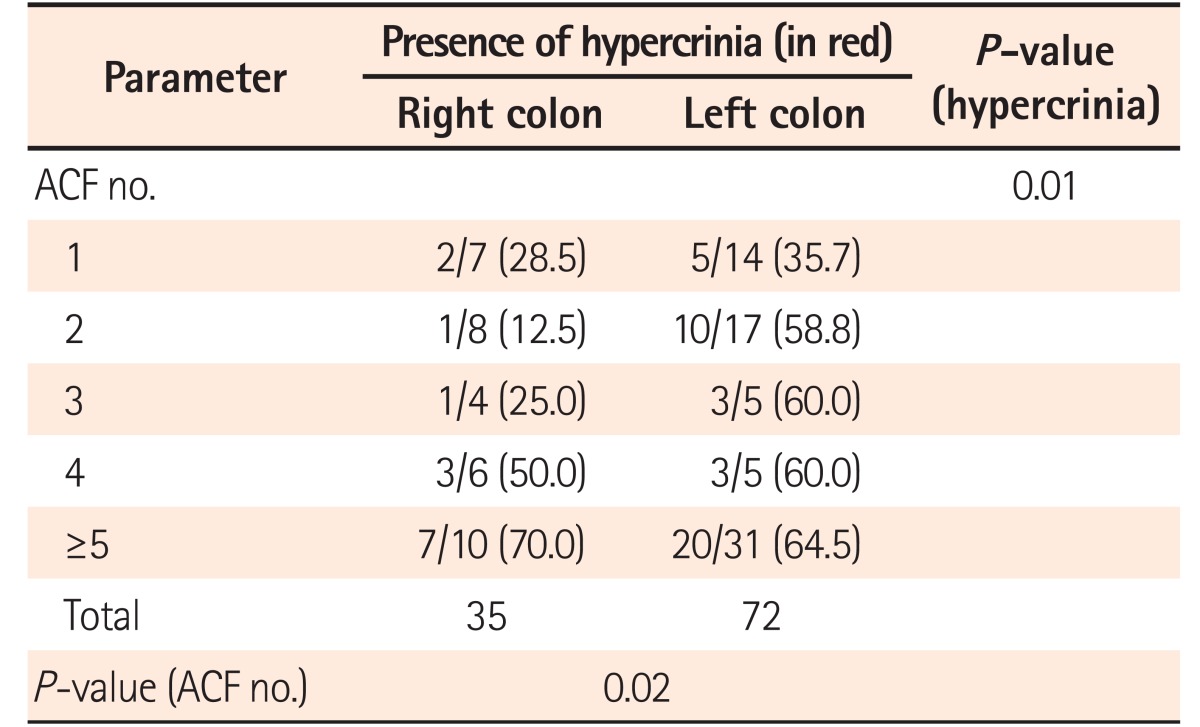
9 We identified a greater number of ACF in the left colon than in the right colon. In colonic mucosa adjacent to CRC, the ACF identified showed increased number, density, size, and crypt multiplicity compared with those of the controls. In comparison to the ACF situated in right colon, the left colonic ACF had more risk factors, such as dysplasia and hypercrinia (increased goblet cell population). Hypercrinia was also more prevalent in colonic mucosa adjacent to foci of CRC and in the ACF with high crypt multiplicity (≥5/ACF). Our 3-tier ACF pit pattern classification showed good correlation with ACF histology and the presence of hypercrinia. The assessment of MSI was performed by the immunohistochemical method, and no statistically significant difference was noted between MMR protein negativity and ACF histology. Across the study groups, although the number of IHC-negative cases (mutated/methylated) was found to be less, there was a slight predominance of the loss of microsatellite markers in ACF associated with CRC (group A). There was significantly greater p53 protein overexpression in both the left-sided ACF and the ACF associated with CRC. The
KRAS gene mutation was found more frequently in the left-sided ACF, one of which showed histological dysplasia, and there was no association between the
KRAS gene mutation and hypercrinia. The
BRAF V600E mutation was not identified in any of the ACF or corresponding tumor samples that were examined, irrespective of laterality.
A previously published study employing confocal chromoendoscopy that was performed on 861 subjects (normal subjects, adenoma cases, and CRC cases), showed an increasing prevalence of ACF in cases of CRC compared to that in the adenomas and in normal colonic mucosa, suggesting that ACF may be an early surrogate microscopic marker for human colon carcinogenesis.
13 In a study published by Roncucci et al.,
14 greater ACF density was found in diseases, such as familial adenomatous polyposis and CRC than in benign colonic diseases, with a positive ACF density gradient from the proximal to the distal colon. Our results were in accordance with these studies. A topographic classification system that can triage the ACF according to their risk of developing CRC in future has been lacking. Based on a large endoscopic study on crypt patterns incorporating normal colonic mucosa, polyps, adenomas, and carcinomas, a system of crypt pattern classification was described by Kudo et al.
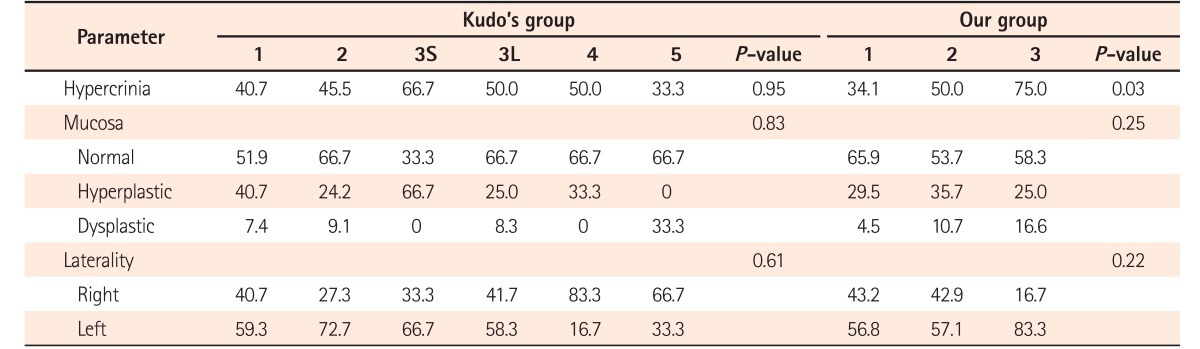
10 According to Kudo's system, the slit-like pattern, gyriform pattern, and non-structural pits were categorized as high-risk crypts, whereas the rounded crypts were the most innocuous. In the study by Roncucci et al.,
15 the round luminal pattern was determined to be the benign type, whereas the slit-like ACF were commonly associated with histologic dysplasia. In our study, the round/oval ACF pit pattern was the most common, whereas the ACF with gyriform and slit-like pit patterns had a more dysplastic histology, thereby qualifying as high-risk ACF subtypes. However, Kudo's classification system was formulated using a large number of crypt patterns and is highly acclaimed; our pit pattern was formulated based on our topographic findings in human mucosal ACF only. Hence, we suggest that our morphological classification method be used solely for triaging ACF and not for other colonic lesions.
Few studies have investigated ACF in human colonic mucosa in terms of MMR protein expression. Although IHC cannot differentiate between a true MMR gene mutation and epigenetic silencing, the utility of IHC to diagnose MSI in CRC has been previously validated by molecular methods. The issue of the interpretation of IHC results as positive or negative has been variously addressed, though predominantly in tumors. To date, no study has examined the expression of MMR proteins in ACF by the IHC method. However, MMR gene status has been studied in ACF by molecular methods.
16 Therefore, our IHC interpretation is subject to validation using molecular methods. While strong diffuse staining of MMR proteins in tumor tissue implies intact MMR gene expression, focal weak staining or the loss of staining in tumor tissue imply mutated MMR genes or the epigenetic silencing of MMR genes.
17 The interpretation of MMR staining in ACF remains unclear. Moreover, there is a wide variation in the results reported in the published literature regarding MMR gene expression. In some studies, loss of expression of these proteins was found in only 2% of tumors,
18,19 whereas others have shown MLH1 inactivation through epigenetic silencing in up to 90% of sporadic MSI+ cancers.
20,21,22 This epigenetic alteration is speculated to be an early event in sporadic CRC that arises through serrated, non-dysplastic adenomas.
22,23 Greenspan et al.
24 identified hMLH1 promoter hypermethylation in 8 of 39 ACF (21%), with no correlation found between hMLH1 promoter hypermethylation and MSI. In this study, we identified MLH1 protein loss in 20% of ACF. In 3 of our ACF, the complete loss of all MMR markers was observed. Although we identified differential MMR protein expression patterns in some ACF, we adopted an all-or-none approach in which ACF with diffuse expression were considered MSS, whereas ACF with partial or no stain expression were considered microsatellite instable lesions. In a study by Cheng and Lai,
16 the authors identified differential MMR gene expression patterns in various ACF found in the colonic mucosa of a single patient. While some of the ACF had MSI, the others showed the MSS pattern. Similarly, no correlation between MMR protein expression and ACF histology was found in the present study.
16
Although the
TP53 mutation has been considered a late event in carcinogenesis,
11 almost 30% of the left-sided ACF had low-grade dysplasia with increased p53 protein expression. Whether all of these dysplastic ACF will eventually progress to adenomas or not, and what is the estimated timeframe of such a progression are not conclusively known. Yamashita et al.
25 observed more frequent
TP53 mutations in left colonic ACF than in right colonic ACF, and increased p53 positivity in left colonic ACF was found in our study as well (60.5% vs. 38.2%,
P=0.03).
11
The serrated pathway is implicated with sessile serrated adenomas and polyps in the right colon and is characterized by a distinct molecular signature involving mutations in mismatch repair (
MMR) genes and in the
BRAF gene. In a recent study published by Inoue et al.,
8 serrated polyps identified in the right colon were associated with ACF found in the vicinity, both of which showed mutation of the
BRAF gene and gene promoter methylation. In a series of 55 cases of ACF, Rosenberg et al.
26 reported
BRAF V600E mutation in 10 of 16 serrated lesions, as compared to that in only 1 of 33 non-serrated lesions (
P=0.001).
KRAS mutation was detected in 3 of 16 serrated ACF, in comparison to that in 14 of 33 non-serrated ACF lesions. The authors also found that 11% of the hyperplastic ACF (5/45) and 25% (1/4) of the dysplastic ACF were the MSI-H type. MSI was not associated with the
BRAF mutation in ACF, indicating that MSI was a late event in the serrated polyp pathway. Beach et al.
27 also showed that
BRAF mutation in ACF was associated with hyperplastic polyposis. In our study, no serrated pattern was noted in any of the ACF identified. None of our ACF showed
BRAF mutation, whereas 3 ACF showed
KRAS mutation. Loss of MMR proteins was not significantly different between the left and right colonic ACF in the index study.
To our knowledge, the presence of hypercrinia in ACF has not been previously described, and the term has mainly been used in relation to crypt changes in IBD, especially CD.
28 Hypercrinia was identified frequently in left colonic ACF in our study. ACF with low-grade dysplasia also showed hypercrinia. These results all suggest that hypercrinia can be used as a histological marker for high-risk ACF. The potential link between excessive mucin production and the pathogenesis of these lesions remain to be investigated further.
Given that ACF-like lesions can be identified in vivo using the magnified chromoendoscopy technique, herein, we aimed to create a study design that would closely mimic a magnified chromoendoscopic view. Based on the current knowledge, chromoendoscopic screening of the whole colon in high-risk subjects is not yet indicated. Therefore, we needed to assess the presence of any topographic, histological, and genetic differences in ACF found in human colectomies belonging to different clinical groups. Based on our findings, it is apparent that there are no indisputable criteria that can accurately differentiate the ACF found in these groups. On multivariate analysis, only p53 protein overexpression was found to be significantly associated with high-risk ACF. The findings may indicate that any ACF in human colon should not be discounted during chromoendoscopy. There are several limitations in this study. First, an elaborative study of genetic change was not performed. We undertook this work as a pilot study and further detailed genetic workups are being performed. Second, we did not validate the MMR protein expression pattern with MMR gene amplification. In conclusion, no significant topographic, histological, and genetic differences exist in ACF identified in different clinical settings. High-risk ACF appear mostly in the left colon and show increased density, crypt multiplicity, hypercrinia, and p53 accumulation. The topographic pit pattern described in this study is simple to remember and can predict a greater likelihood of histological hypercrinia and dysplasia.
SUPPORTING INFORMATION
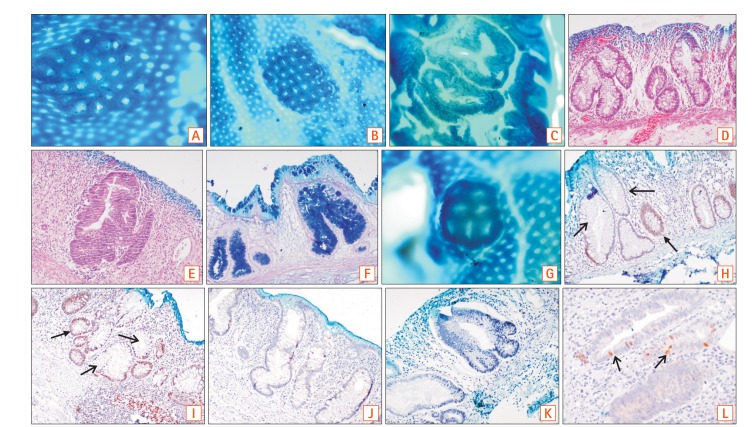
1. Aberrant crypt foci (ACF) are a cluster of dilated and occasionally distorted crypts in comparison to the surrounding area, with a large pericryptal zone. The thick epithelial lining stains darker with methylene blue as compared to normal crypts. The luminal opening is 2 to 3 times larger than normal and the crypt pit pattern can be variable. In some cases, ACF were seen as a single aberrant crypt and not as a cluster.
2. ACF density refers to the number of ACF per square centimeter of mucosal surface.
3. Elevated ACF implies that the luminal openings lie above the level of the adjacent normal mucosa when viewed end-on as in magnified chromoendoscopy or as in our case under 40× magnification microscopy. In contrast, flat ACF have luminal openings at the same level as that of adjacent normal mucosa.
4. Hyperplastic ACF refers to ACF with hyperplastic epithelial lining and no dysplasia on histology.
5. Dysplastic ACF (microadenoma) show nuclear stratification, enlargement, hyperchromasia, loss of polarity, prominent nucleoli, and irregular outline on histology. We divided dysplastic ACF into low-grade and high-grade types based on the standard definitions of dysplasia used in pathology.
6. Hypercrinia refers to an increased number of goblet cells with near extinction of absorptive cells and increased mucin production on light microscopy.
7. Kudo's classification is a 5-tier endoscopic classification system (1, roundish; 2, stellar or papillary; 3S, small-roundish or tubular; 3L, large-roundish or tubular; 4, branch-like or gyrus; and 5, non-structured pits) to identify the various morphological appearances of colonic crypts described with magnifying endoscopes after methylene blue staining of the mucosa.10
8. Our proposed topographic classification system has been devised to identify ACF after methylene blue staining and can be applied to chromoendoscopy as well. It is a 3-tier system that describes round, slit-like, and gyriform pits (
Fig. 1A-C).